pelt
stephens999
2018-10-23
Last updated: 2018-10-23
workflowr checks: (Click a bullet for more information)-
✔ R Markdown file: up-to-date
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
-
✔ Environment: empty
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
-
✔ Seed:
set.seed(20180414)The command
set.seed(20180414)was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible. -
✔ Session information: recorded
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
-
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.✔ Repository version: 222fa03
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can usewflow_publishorwflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.Ignored files: Ignored: .DS_Store Ignored: .Rhistory Ignored: .Rproj.user/ Ignored: analysis/.Rhistory Untracked files: Untracked: analysis/cp_convergence.Rmd Untracked: analysis/cp_init_tf.Rmd Untracked: analysis/null.Rmd Untracked: analysis/test.Rmd Untracked: data/geneMatrix.tsv Untracked: data/liter_data_4_summarize_ld_1_lm_less_3.rds Untracked: data/meta.tsv Untracked: docs/figure/cp_convergence.Rmd/ Untracked: docs/figure/test.Rmd/
Expand here to see past versions:
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 222fa03 | stephens999 | 2018-10-23 | workflowr::wflow_publish(“analysis/pelt.Rmd”) |
Introduction
The idea here is to check into why the default method for cpt.mean function did not perfom well on some examples.
library(changepoint)Loading required package: zoo
Attaching package: 'zoo'The following objects are masked from 'package:base':
as.Date, as.Date.numericSuccessfully loaded changepoint package version 2.2.2
NOTE: Predefined penalty values changed in version 2.2. Previous penalty values with a postfix 1 i.e. SIC1 are now without i.e. SIC and previous penalties without a postfix i.e. SIC are now with a postfix 0 i.e. SIC0. See NEWS and help files for further details.set.seed(51)
true_mean = rep(c(-0.2,0.1,1,-0.5,0.2,-0.5,0.1,-0.2),c(137,87,17,49,29,52,87,42))
genomdat = list(x = rnorm(500, sd=0.2) + true_mean, true_mean=true_mean)The cpt.mean default does not find any changepoints:
genomdat.cp = cpt.mean(genomdat$x,method="PELT")
plot(genomdat.cp)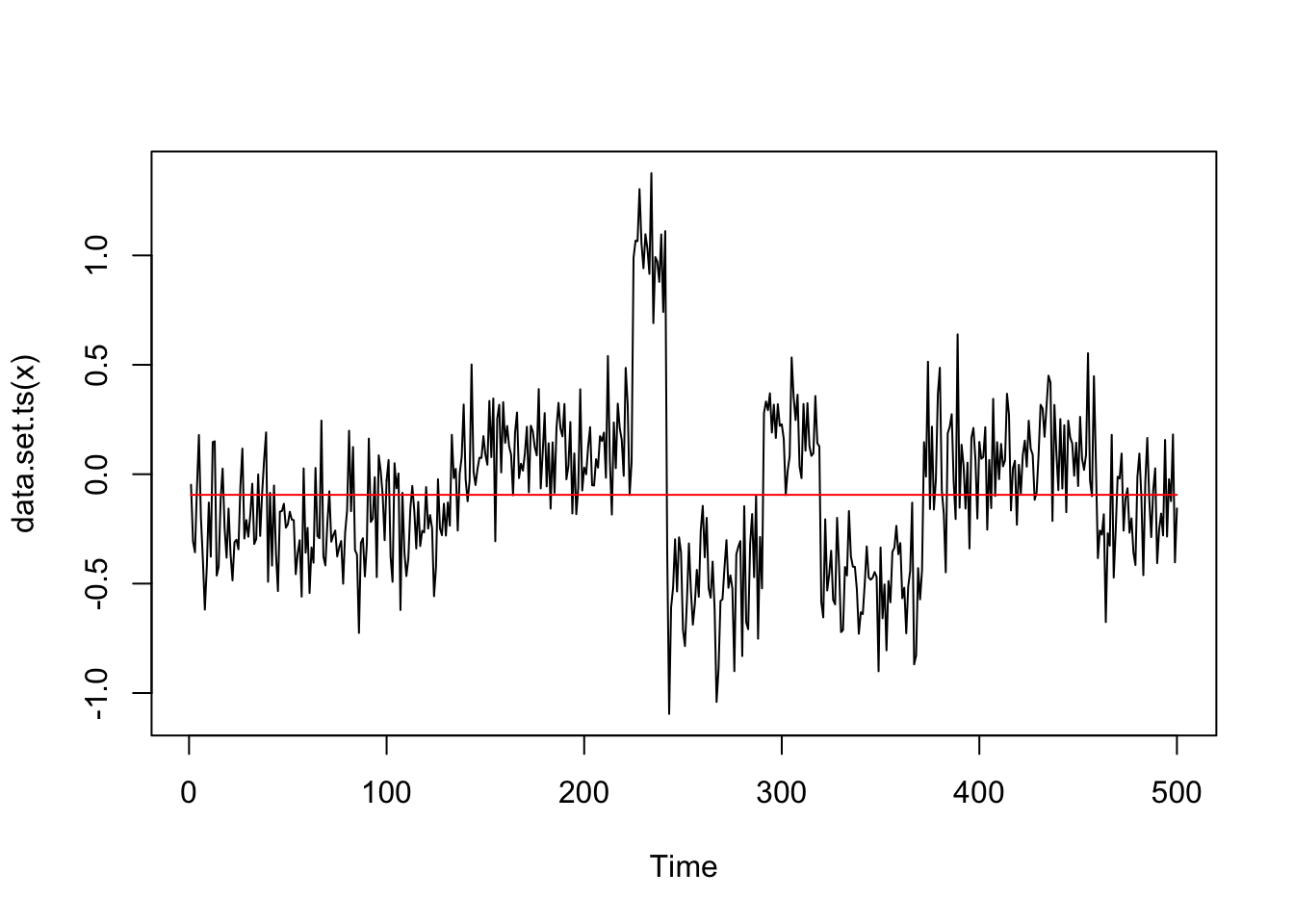
Interestingly if we just multiply the data by 10 we find many changepoints.
genomdat.cp = cpt.mean(10*genomdat$x,method="PELT")
plot(genomdat.cp)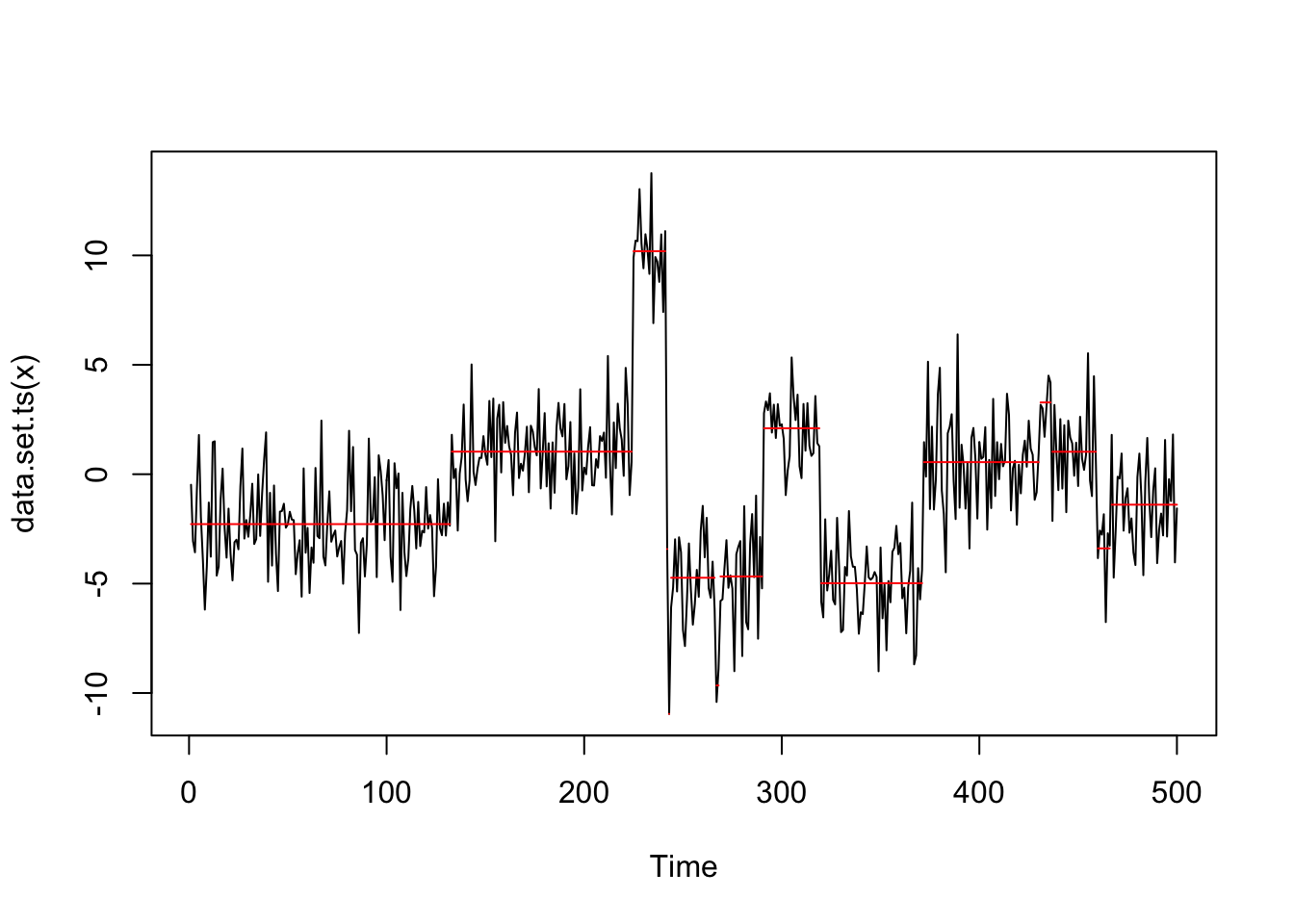
And if we multiply by 100 we find many, many changepoints
genomdat.cp = cpt.mean(100*genomdat$x,method="PELT")
plot(genomdat.cp)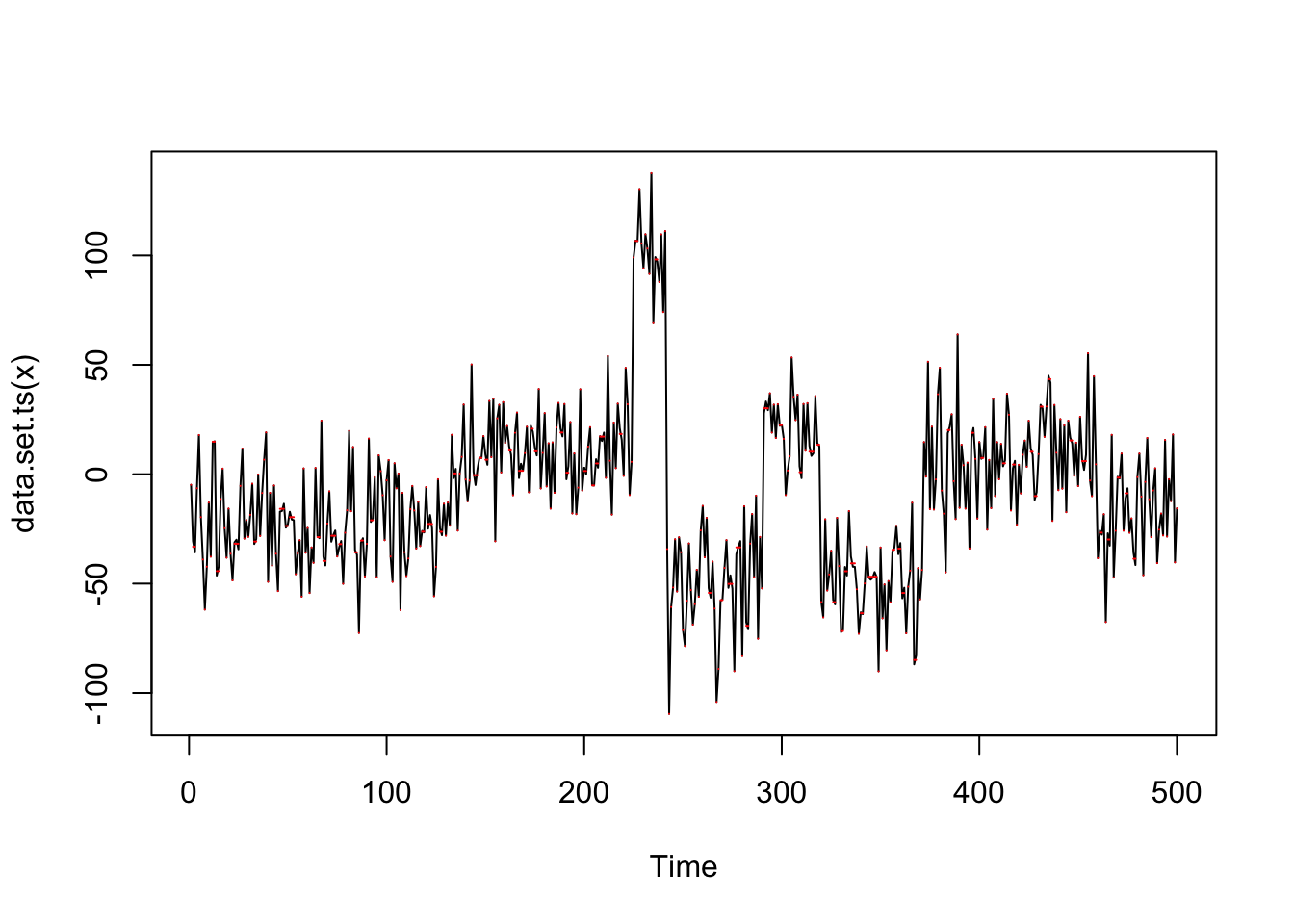
I suspect that the cost function may implicitly assume residual variance is 1; I opened an issue on the checkpoint github page.
Session information
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: OS X El Capitan 10.11.6
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] changepoint_2.2.2 zoo_1.8-4
loaded via a namespace (and not attached):
[1] workflowr_1.1.1 Rcpp_0.12.19 lattice_0.20-35
[4] digest_0.6.18 rprojroot_1.3-2 R.methodsS3_1.7.1
[7] grid_3.5.1 backports_1.1.2 magrittr_1.5
[10] git2r_0.23.0 evaluate_0.12 stringi_1.2.4
[13] whisker_0.3-2 R.oo_1.22.0 R.utils_2.7.0
[16] rmarkdown_1.10 tools_3.5.1 stringr_1.3.1
[19] yaml_2.2.0 compiler_3.5.1 htmltools_0.3.6
[22] knitr_1.20 This reproducible R Markdown analysis was created with workflowr 1.1.1