Real Data with Simulated Signals: Part IV
Lei Sun
2017-05-22
Last updated: 2017-12-21
Code version: 6e42447
Introduction
We are comparing our method with ASH and SVA.
library(ashr)
library(edgeR)
library(limma)
library(seqgendiff)
library(sva)
library(pROC)source("../code/gdash.R")Artefactual effects \(\pi_0\delta_0 + \left(1 - \pi_0\right)N\left(0, \sigma^2\right)\) are added to the real GTEx data.
mat = read.csv("../data/liver.csv")We are using \(10K\) genes, and \(100\) independent simulation trials.
ngene = 1e4
N = 100Setting 1: \(2\) v \(2\), \(\pi_0 = 0.9\), \(\sigma^2 = 1\)
nsamp = 4
pi0 = 0.9
sd = 1
set.seed(777)
system.time(gdash.comp <- N_simulations(N, mat, nsamp, ngene, pi0, sd))Coefficients not estimable: 3 Warning: Partial NA coefficients for 10000 probe(s)Warning in ebayes(fit = fit, proportion = proportion, stdev.coef.lim =
stdev.coef.lim, : Estimation of var.prior failed - set to default valueWarning: Zero sample variances detected, have been offsetCoefficients not estimable: 3 Warning: Partial NA coefficients for 10000 probe(s)Warning in ebayes(fit = fit, proportion = proportion, stdev.coef.lim =
stdev.coef.lim, : Estimation of var.prior failed - set to default valueCoefficients not estimable: 3 Warning: Partial NA coefficients for 10000 probe(s)
Warning: Estimation of var.prior failed - set to default value user system elapsed
4082.811 380.118 1649.864 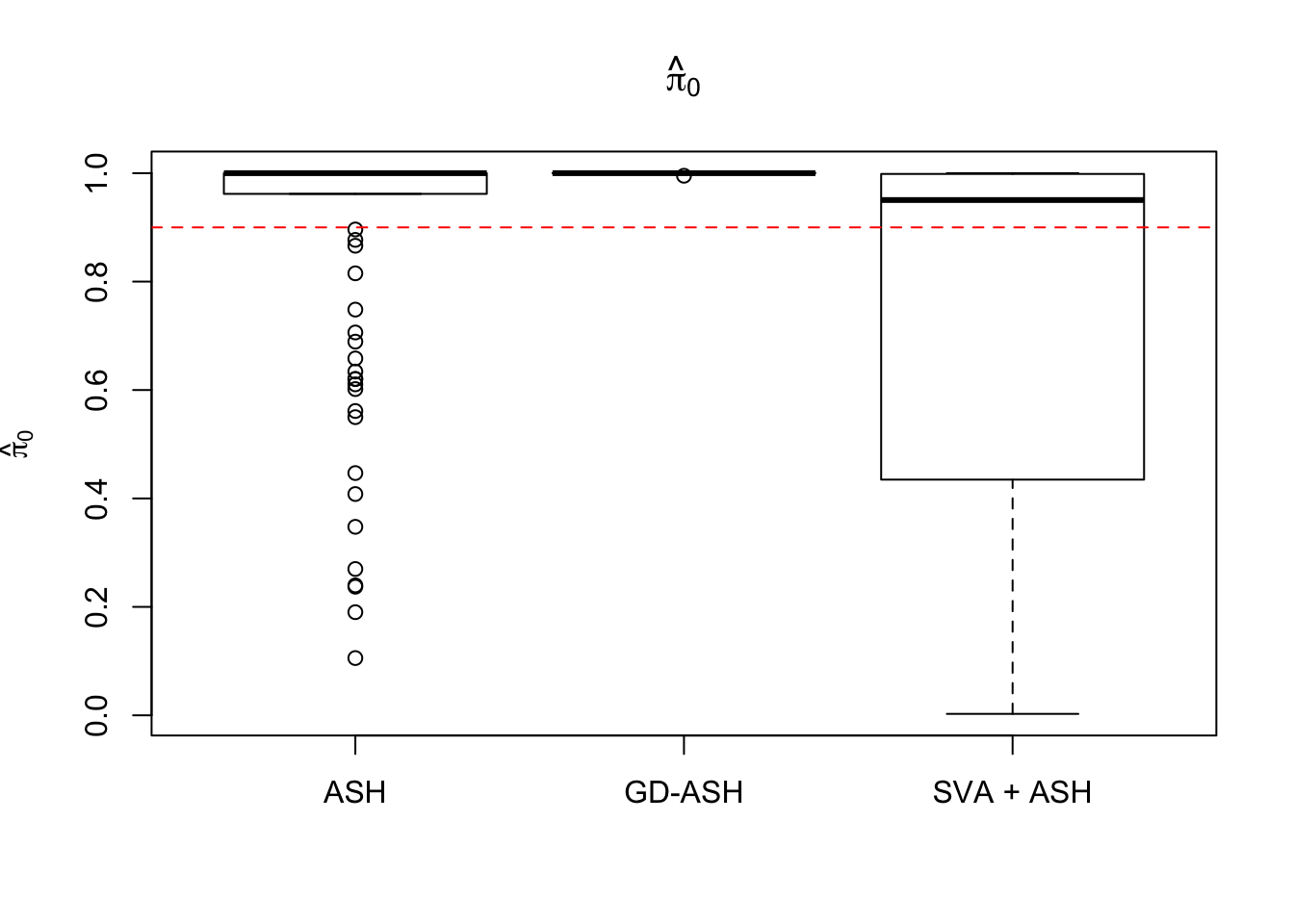
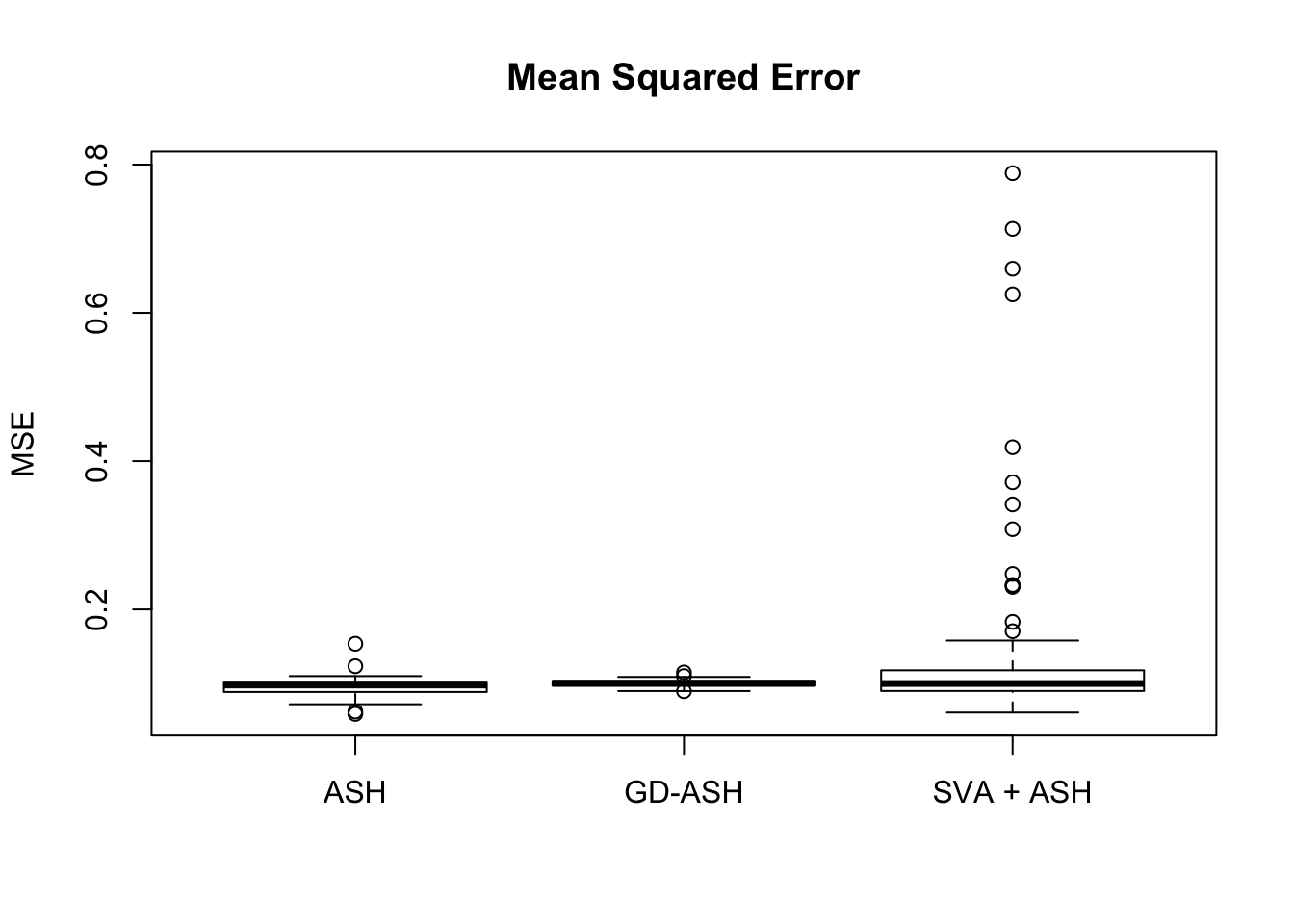
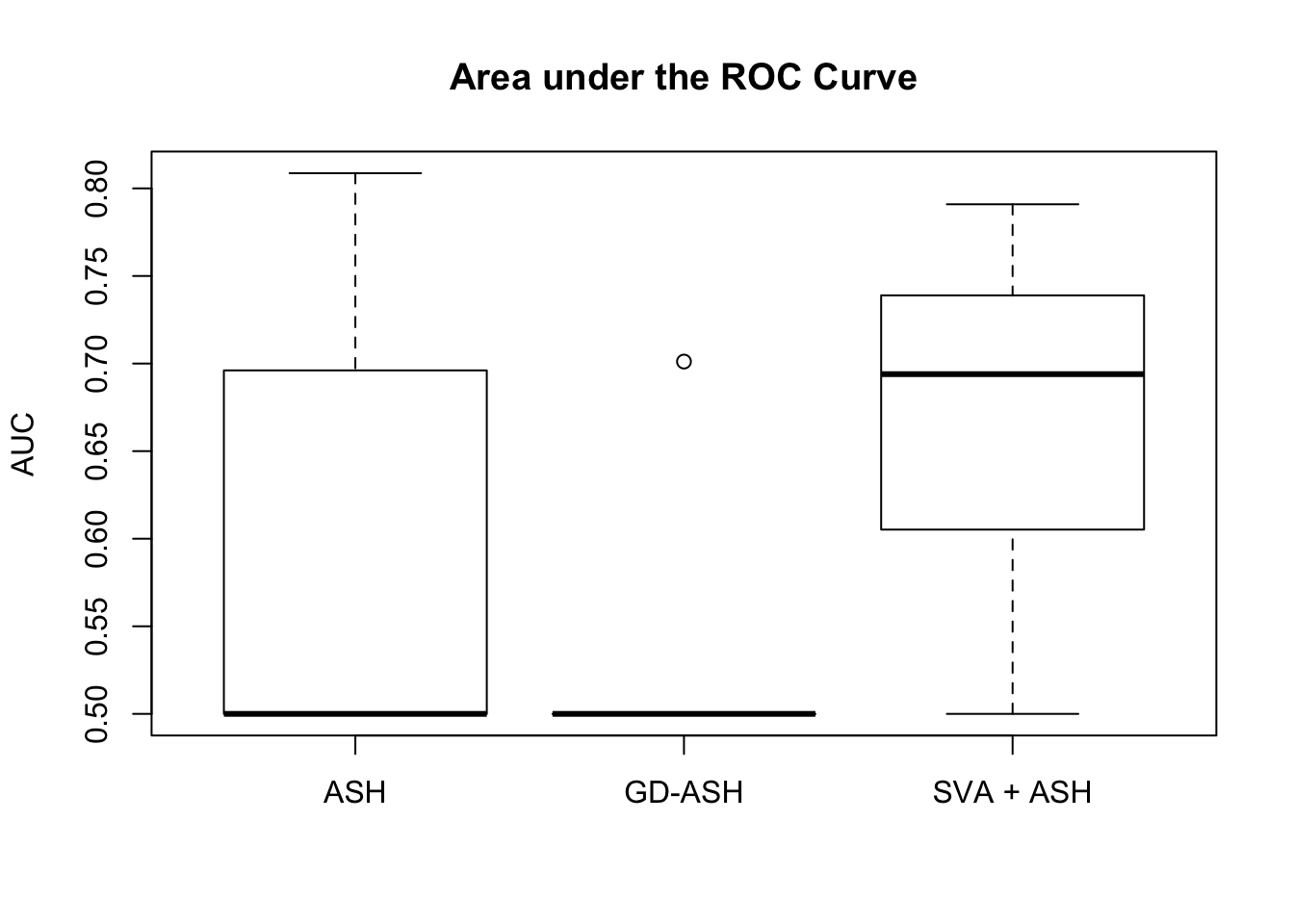
Setting 2: \(3\) v \(3\), \(\pi_0 = 0.9\), \(\sigma^2 = 1\)
nsamp = 6
pi0 = 0.9
sd = 1
set.seed(777)
system.time(gdash.comp <- N_simulations(N, mat, nsamp, ngene, pi0, sd)) user system elapsed
4235.948 319.028 1655.836 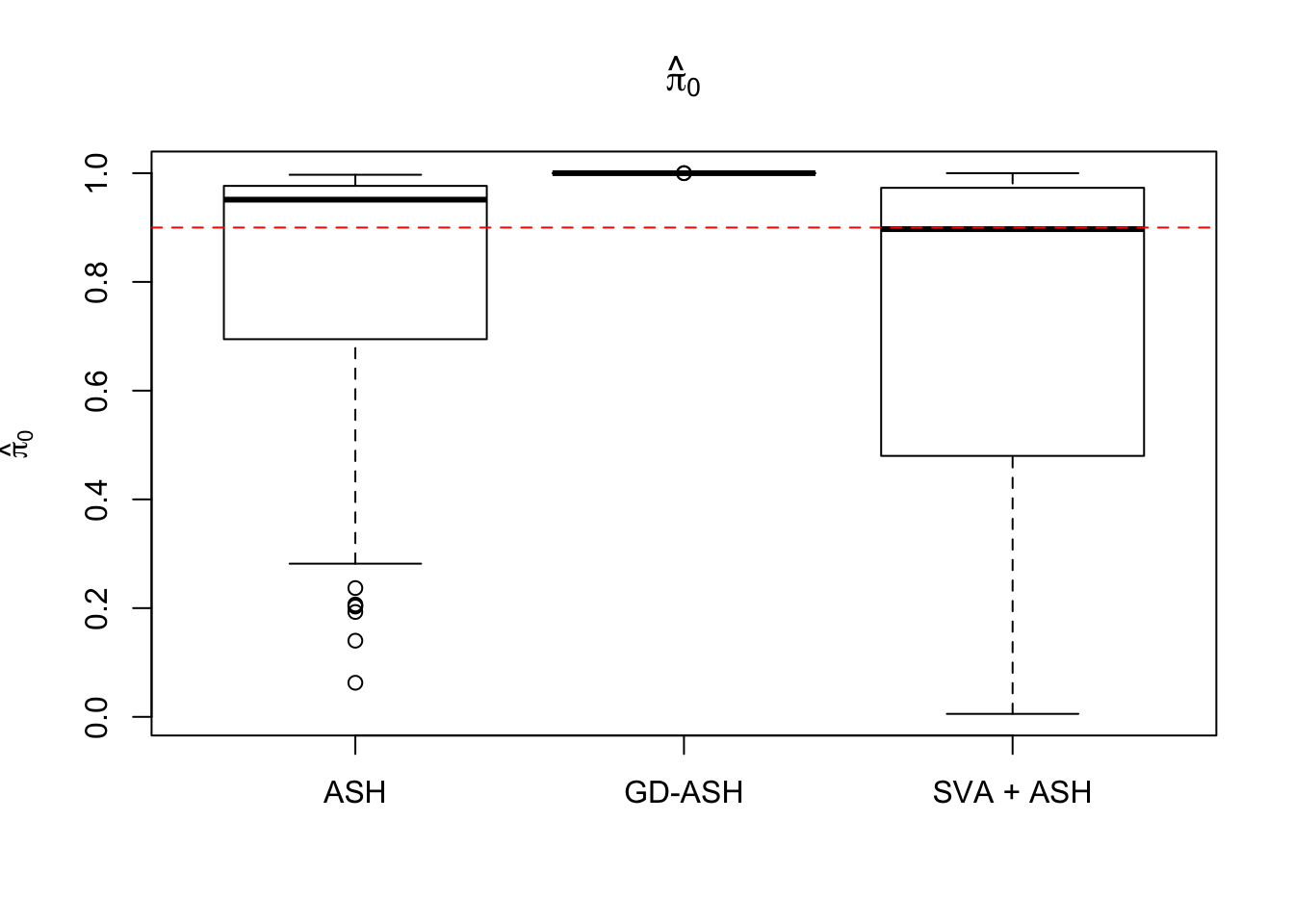
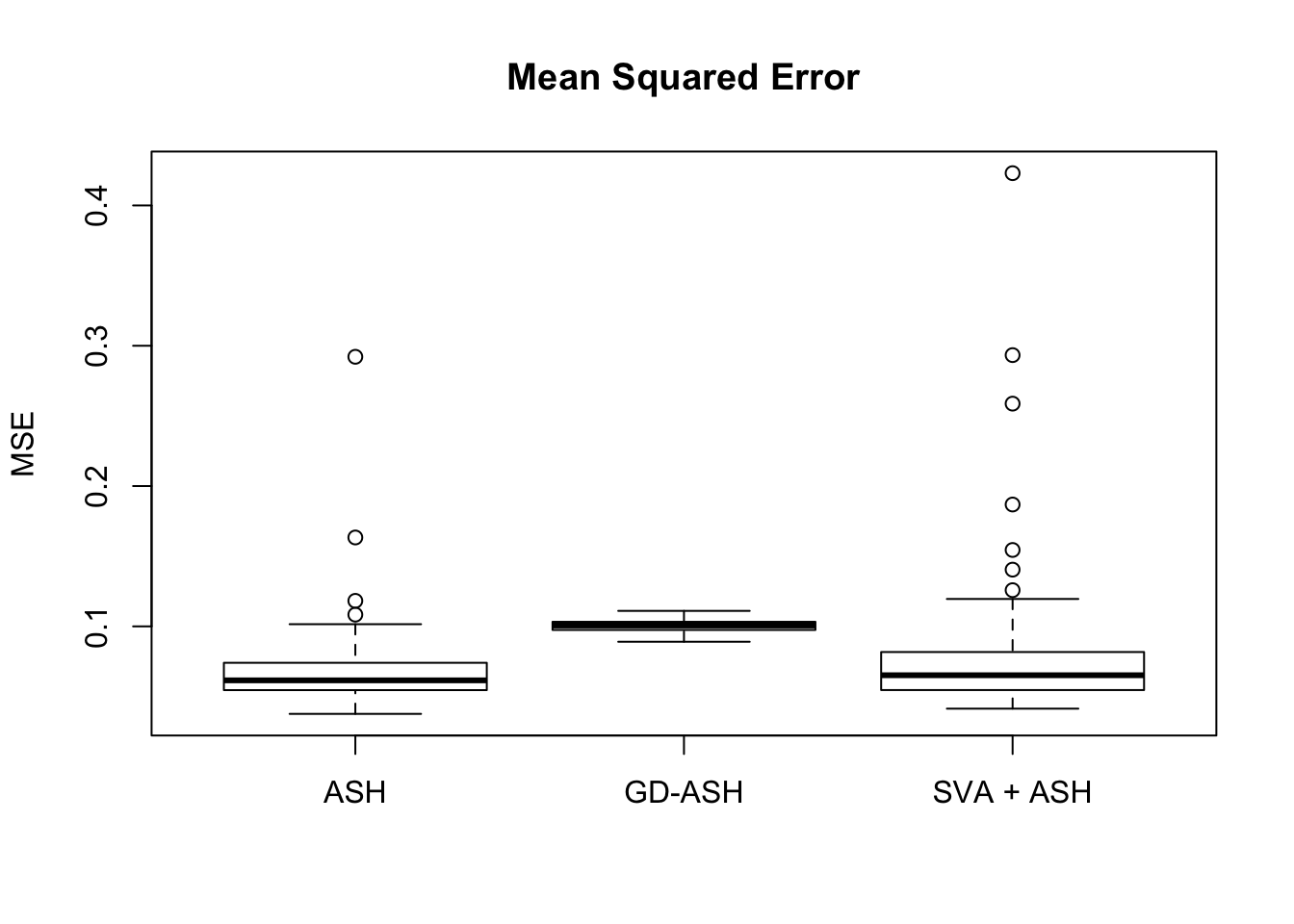
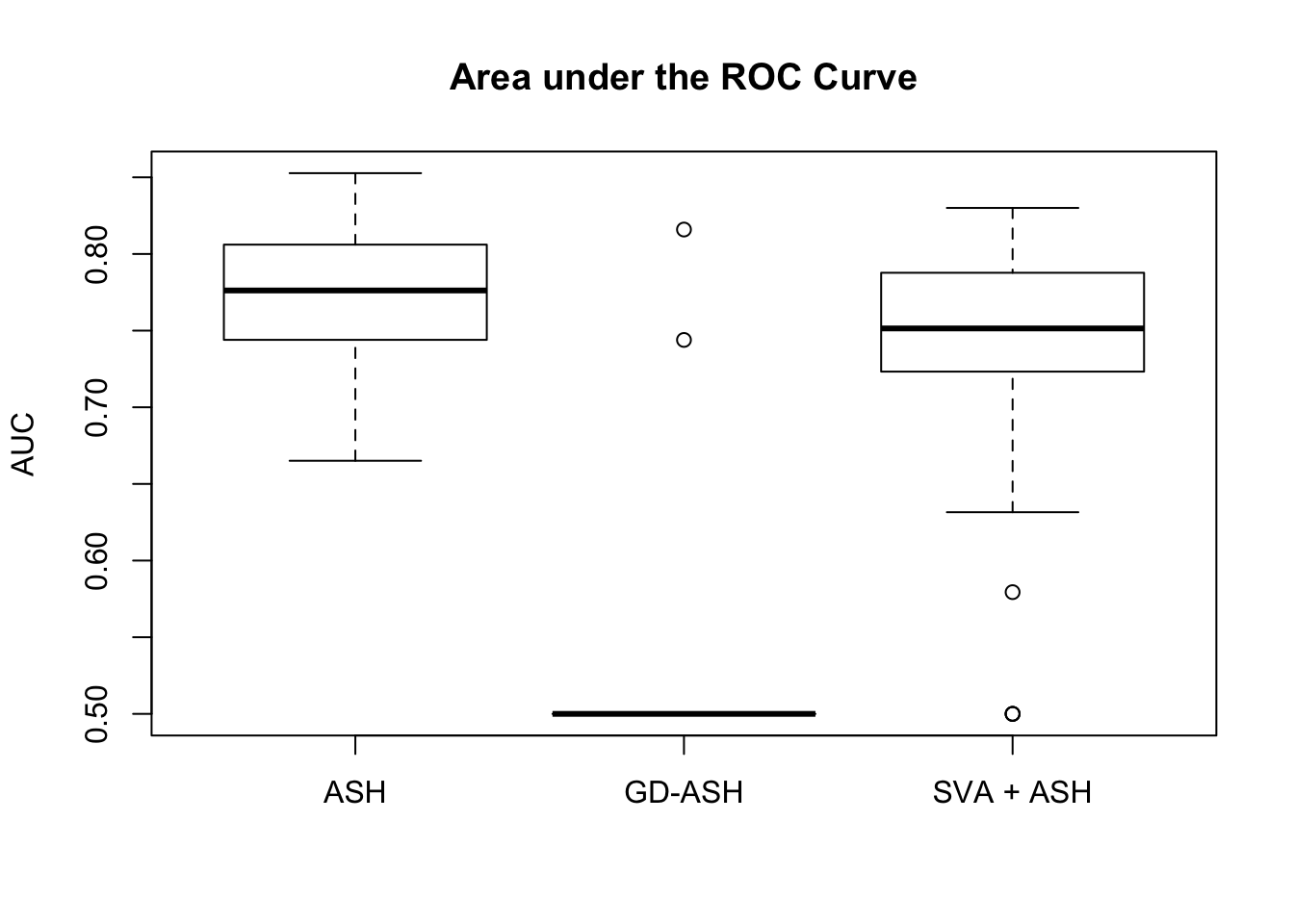
Session information
sessionInfo()R version 3.4.3 (2017-11-30)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS High Sierra 10.13.2
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
loaded via a namespace (and not attached):
[1] compiler_3.4.3 backports_1.1.2 magrittr_1.5 rprojroot_1.3-1
[5] tools_3.4.3 htmltools_0.3.6 yaml_2.1.16 Rcpp_0.12.14
[9] stringi_1.1.6 rmarkdown_1.8 knitr_1.17 git2r_0.20.0
[13] stringr_1.2.0 digest_0.6.13 evaluate_0.10.1This R Markdown site was created with workflowr